NVIDIA has unveiled Proteina-Complexa, an innovative generative model designed to create de novo protein binders and enzymes. This development addresses the intricate challenges involved in designing protein binders, which are essential for therapies and catalysts. Achieving strong and specific binding to target proteins or small molecules necessitates a meticulous optimization of interactions, given the vast array of amino acid sequence permutations and 3D protein structures.
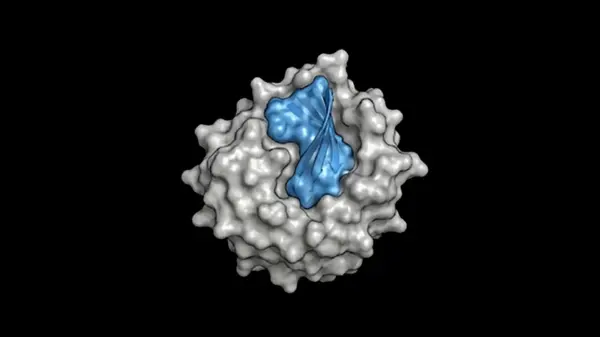
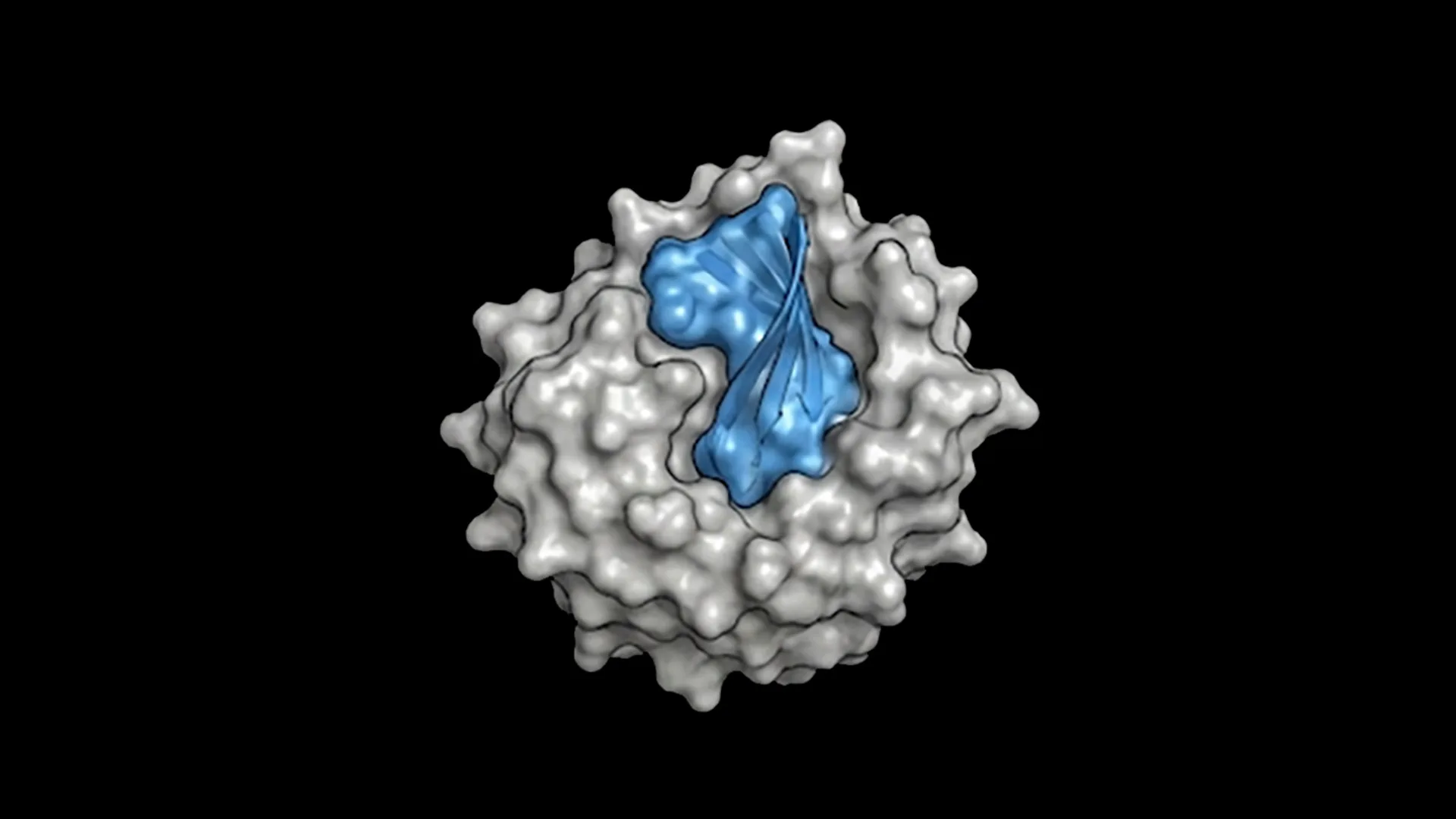
Proteina-Complexa leverages a partially latent flow-matching framework built atop the La-Proteina model. This advanced approach allows for the simultaneous generation of both the fully atomistic structure of a binder—including its protein backbone and side-chains—and the corresponding amino acid sequence, termed co-design. By modeling backbone alpha carbon atoms in 3D Cartesian space and compressing all other atoms into a learned latent space through an autoencoder, the system strikes a balance between atomic fidelity and computational efficiency.
Traditionally, workflows for binder design have been fragmented, often requiring separate models for backbone and sequence generation. In contrast, Proteina-Complexa’s co-design capabilities facilitate reasoning at an atomistic level, ensuring that chemical identities and 3D geometries are tightly coupled. This integrated generation process enhances the design of precise, high-affinity interfaces, which are optimized for folding and synthesis.
To support its generative model, Proteina-Complexa was trained using over 1 million curated high-quality experimental and predicted structures from notable databases, including the Protein Data Bank (PDB) and AlphaFold Protein Structure Database. The model introduces a unified approach that combines generative techniques with inference-time compute scaling, allowing for iterative design optimization during inference. Reasoning search algorithms evaluate and refine candidates at intermediate steps, enhancing both computational efficiency and the quality of generated binders.
Proteina-Complexa can generate protein binders for various applications, including designing binders for protein targets in oncology, immunology, and neurology, as well as for small molecule targets and enzyme design. Experimental validation has been a critical component of its development, with extensive collaborations yielding results. For instance, successful binder designs have been validated through partnerships with organizations such as Manifold Bio, Novo Nordisk, and Duke University, showcasing the model’s versatility.
In one notable experiment, Proteina-Complexa produced binders against the Activin Receptor Type-2A, a target linked to muscle-wasting disorders, achieving affinities in the nano- and picomolar range. The system has also demonstrated its capability to bind to sugars on red blood cells, a task typically fraught with challenges due to the polar nature of carbohydrates. This was validated by identifying four candidates that exhibited strong agglutination signals, surpassing the efficacy of existing natural protein-based alternatives.
The validation process involved generating tens of millions of initial in silico candidates, subsequently filtering to around 1 million for experimental testing against 133 distinct protein targets. Large-scale experiments utilizing multiplexed phage screening technology formed one of the largest binder design benchmarks to date, with quantitative binding kinetics assessed using techniques such as surface plasmon resonance and western blotting.
Looking ahead, Proteina-Complexa provides users with the tools to customize and adapt its capabilities for various research and development needs. The model’s source code, trained checkpoints, and datasets are available to facilitate further exploration into protein-based therapeutics, catalysts, and biosensors. As researchers and developers engage with Proteina-Complexa, the potential for groundbreaking advancements in the field of protein design increases, paving the way for innovative therapeutic solutions.
See also Sam Altman Praises ChatGPT for Improved Em Dash Handling
Sam Altman Praises ChatGPT for Improved Em Dash Handling AI Country Song Fails to Top Billboard Chart Amid Viral Buzz
AI Country Song Fails to Top Billboard Chart Amid Viral Buzz GPT-5.1 and Claude 4.5 Sonnet Personality Showdown: A Comprehensive Test
GPT-5.1 and Claude 4.5 Sonnet Personality Showdown: A Comprehensive Test Rethink Your Presentations with OnlyOffice: A Free PowerPoint Alternative
Rethink Your Presentations with OnlyOffice: A Free PowerPoint Alternative OpenAI Enhances ChatGPT with Em-Dash Personalization Feature
OpenAI Enhances ChatGPT with Em-Dash Personalization Feature