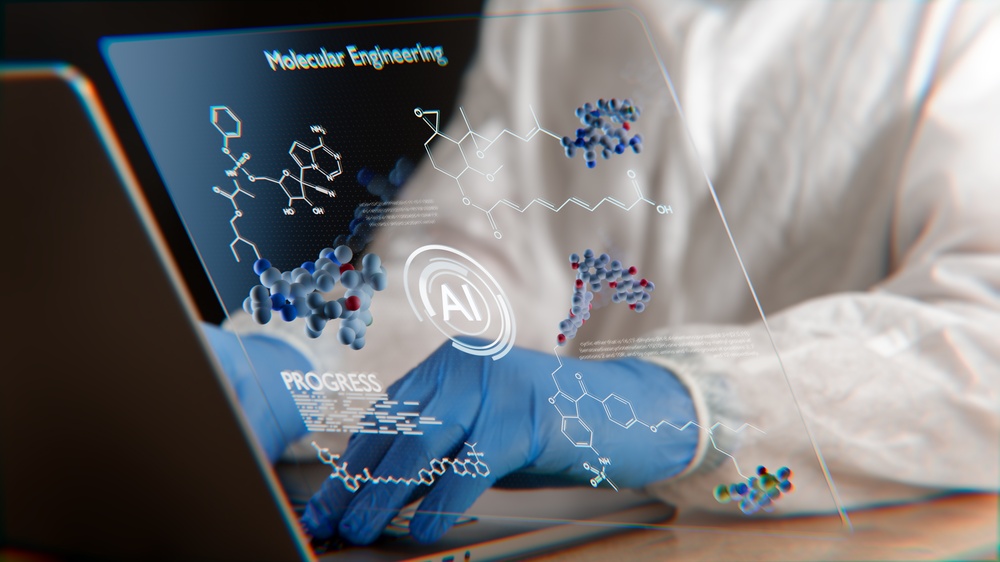
Scientists at the University of Virginia School of Medicine have developed a new method of drug design that leverages artificial intelligence diffusion models to track and adapt to protein movements. The innovative suite of tools, named YuelDesign, YuelPocket, and YuelBond, aims to enhance the efficiency of the drug development process while also identifying new ways to improve the effectiveness of existing medications.
The financial burden of developing a new drug is substantial, with costs ranging from several hundred million dollars to over $2.6 billion and continuing to escalate. Furthermore, around 90% of new drugs fail to pass human testing trials, largely due to unpredictable interactions between drug molecules and their biological targets, leading to treatments that may have little or even detrimental effects.
The challenge lies in the nature of proteins, which frequently undergo “induced fits”—a phenomenon where they change shape upon binding with a drug. Traditionally, predicting these changes using computerized models has proven difficult. However, the advanced AI diffusion models developed in this project are capable of generating the structure of the protein pocket and identifying small molecules that can effectively bind to it, thereby facilitating more adaptive design.
“Think of it this way: Other methods try to design a key for a lock that’s sitting perfectly still, but in your body, that lock is constantly jiggling and changing shape,” explained Nikolay V. Dokholyan, PhD, from UVA’s Department of Neurology. “Our AI designs the key while the lock is moving, so the fit is much more realistic.” This innovative approach holds promise for patients with cancer, neurological disorders, and various other conditions that desperately require improved drug therapies targeting these dynamic proteins.
The three tools work in unison to optimize drug design. YuelDesign employs the diffusion model to create customized shapes for drug molecules. In parallel, YuelPocket utilizes graph neural networks to accurately identify binding sites on proteins, while YuelBond ensures the precision of chemical bonds within the designed molecules.
“Most existing AI tools treat the protein as a frozen statue, but that’s not how biology works,” stated researcher Dr. Jian Wang. “Our approach lets the protein and the drug candidate evolve together during the design process, just as they would in the body.” Wang noted that during their research, when designing molecules for a well-known cancer-related protein called CDK2, only YuelDesign was able to capture the critical structural changes that occur when a drug binds.
The research team is optimistic that their AI tools will transform drug development into a more cost-effective and timely process, with aspirations to democratize drug discovery. “Our ultimate goal is to make drug discovery faster, cheaper and more likely to succeed, so that promising treatments can reach patients sooner,” Dokholyan remarked. He emphasized their commitment to the scientific community by making all of their tools freely accessible, encouraging researchers around the globe to utilize them in addressing significant medical challenges.
This development reflects a growing trend of employing AI in pharmaceutical research, potentially revolutionizing how drugs are designed and brought to market. As researchers continue to refine these AI methodologies, the hope is that they can significantly reduce the high failure rates in drug development and expedite the delivery of effective treatments to patients in urgent need.
See also Sam Altman Praises ChatGPT for Improved Em Dash Handling
Sam Altman Praises ChatGPT for Improved Em Dash Handling AI Country Song Fails to Top Billboard Chart Amid Viral Buzz
AI Country Song Fails to Top Billboard Chart Amid Viral Buzz GPT-5.1 and Claude 4.5 Sonnet Personality Showdown: A Comprehensive Test
GPT-5.1 and Claude 4.5 Sonnet Personality Showdown: A Comprehensive Test Rethink Your Presentations with OnlyOffice: A Free PowerPoint Alternative
Rethink Your Presentations with OnlyOffice: A Free PowerPoint Alternative OpenAI Enhances ChatGPT with Em-Dash Personalization Feature
OpenAI Enhances ChatGPT with Em-Dash Personalization Feature